FISH stands for fluorescence in situ hybridization testing. In an extremely small nutshell, the essential idea for the testing is that DNA probes, matching the sequence of a specific genetic abnormality being tested are added to a patient's cells or tissue. If the abnormality is present, the probes will bind and the attached fluor will fluoresce. There are many different variations and specifics that I wouldn't be able to do justice to, so will leave it at that simplistic explanation. With improving technology, decreasing costs, and more standard charging, this testing is becoming more and more common in labs across the country, though many still send this testing out to a reference lab. In this post, I'll discuss how I built this for one lab that performed the testing in-house. While there isn't a specific 'cytogenetics' module per se in Beaker (yet), Beaker and Epic as a whole, are incredibly powerful tools with an impressive level of customization.
Ordering FISH in Beaker
Let's start from the top and look at the ordering of FISH. Prior to Beaker, the lab that I built this out in used a paper requisition that allowed physicians to not only order full panels (Acute Myeloid Leukemia panel (AML), Myelodysplastic Syndrome (MDS), etc...) as well as individual probes within the panels (MECOM/RPN1, BCR/ABL + ASS, PML/RARA, etc...). We first needed a way to accomodate this - the ability to order full panels or mix match specific probes. To do this, we used a generic FISH Testing order that had a variety of cascading order questions. The questions would ask if the provider wanted a full panel or specific probes, for each of our panels. If they chose specific probes, the question would cascade to offer a Yes/No for each specific probe. It looked something like this: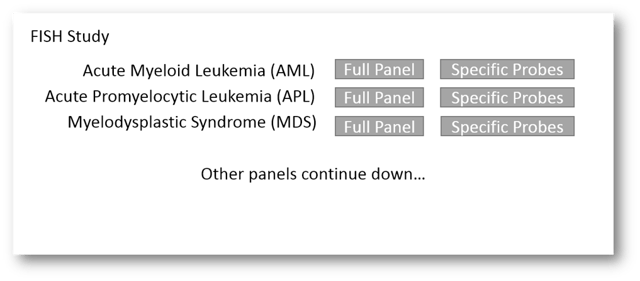 
If the provider chose "Full Panel", no additional order questions would cascade. If, on the other hand, the provider chose "Specific Probes", order questions would cascade asking for the Yes/No for each probe in that panel. That would look something like this:
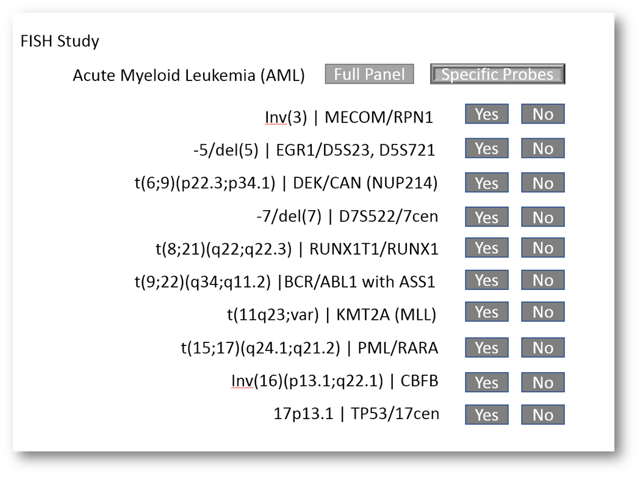
This let the ordering provider specifically choose the probes they were ordering. We had a few additional information questions asked at ordering such as disease status and clinical indication. Your folks will likely have a few that they'd like to ask at ordering as well.
FISH Case Building in Beaker
I did not build anything out of the ordinary for the collection of the specimen so will skip over that and jump to case building. I used one giant protocol that had every possible probe listed in it, to get applied based on rules. To choose the appropriate probes, I built out rules that would look to the answers provided via the order questions. These were tricky as some probes were in multiple panels and could also be ordered individually. So my rules would have logic that looked something like this: 
You'd build out rules for each probe your lab performs. Be sure to include logic for all panels that include the probe in addition to the the probe-specific question.
FISH Task Completion/Slide Creation in Beaker
While we gave a lot of power to ordering providers, it was important that we still give the lab final say on which probes were being tested. So, lab staff would review the tasks that defaulted in, based on the order questions, and make adjustments as needed. We did not put any tracking in the system for the processing, harvesting, etc... work. There are some tools to make that happen, though, including using follow-ups and/or tracking.Resulting FISH in Beaker
We next took advantage of the really cool feature that adds components to a test based on the tasks that are completed. This let us *only* have discrete components listed for the probes that were actually performed. So, if the ordering provider indicated they wanted to test MECOM/RPN1 (by either the probe-specific order question or the panel question), it would fire in a MECOM/RPN1 task. If the lab agreed and completed the task, the result components that went along with this probe would fire in, and importantly, not before. This would be the case for all of your probes. We had the following components per probe: 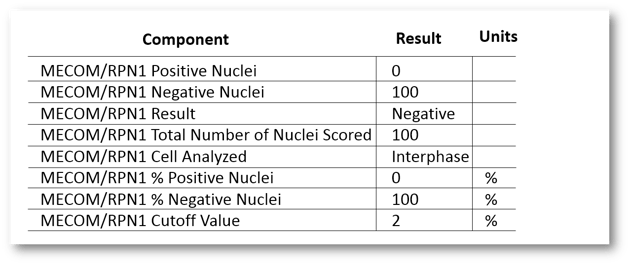
The tech would only manually result the first two components (orange below). The rest would automatically populate via equations or mnemonics (green below).
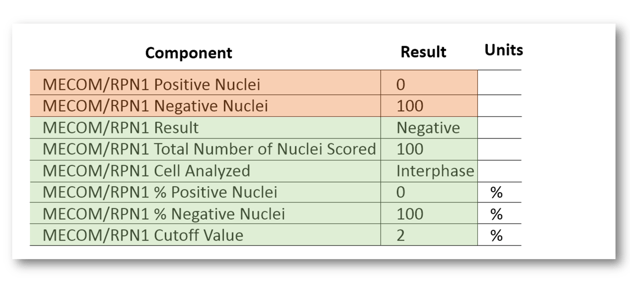
The "Result" component would populate using an equation that compared the % positive nuclei with the cutoff value. We also wanted to match existing reports, which summarized the data in a specific textual format. So, after a value was populated for the "Result" component, I fired a mnemonic that would pull the discrete results under an appropriately formatted text header in another rich text component. That looked something like this:

This would have repeated for each probe that was being tested, automatically of course. In addition to this, we also fired in the probe description into another rich text component. That is pretty straightforward so I won't discuss that here.
